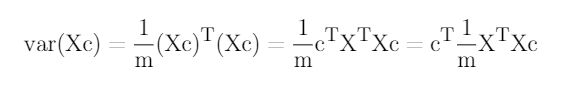This Fall I had the opportunity to advise a student’s senior project! She wanted to study “dimension reduction” which, broadly speaking, consists of taking a large set of data and making it smaller, in various clever ways. I think this could be added to a linear algebra class for a nice real life application.
I was excited to do this because I’ve always wanted to learn a little about data science, but also data science is especially good to know for a mathematician who does not have a tenure track position. To finalize my understanding of what we did I want to try to explain it here. The first thing I learned is that data scientists, just like statisticians, seem to use unnecessary language so I will do my best to write what math people might say instead of what a data scientist would say.
Data is sought after and vacuumed up at alarming rates nowadays. Data can really be any piece of information about something. We want to consider a bunch of data, i.e. pieces of information, about a bunch of things, all at once. One very efficient way to do this is to use matrices. For example, we might ask 2 people (things), for 5 pieces of information:
Person A:
– 6 feet tall
– 190 pounds
– 33 years old
– Likes pineapple on pizza
– Single
Person B:
– 5 feet 4 inches tall
– 140 pounds
– 32 years old
– Does not like pineapple on pizza
– Not Single
We can then write the data in a matrix:

Excuse the crude paint created pictures, but I’ve found this to be much faster than anything else!
There are two data points and in general the number of rows of the matrix is the number of data points. Meanwhile there are 5 “features” that encompass each data point, but really this is the number of columns. Oddly, the number of features is also called the dimension of the data set, even though dimension of a matrix already means the number of rows times the number of columns. To summarize: we can either say there are 5 columns, 5 dimensions, or 5 features to this data set, and that there are 2 data points or 2 rows.
With this mini example the term “dimension reduction” may seem obvious. Dimension reduction involves taking a large set of data represented as a matrix with many columns, and transforming it into a matrix with fewer columns. Of course, there are many ways to make a bigger matrix smaller, but we want to do it in a useful way!
When you google “Principal Component Analysis” or “PCA” you find that it’s a pretty simple way of reducing the dimension of a matrix. However, it took me a while to understand why it was a good way to reduce dimension, so that’s what I want to explain.
To this end let’s just think about two dimensional data, and let’s think about how we could reduce it to one dimension in a smart way. Since we are thinking about two dimensional data this means the data can be written as a matrix with two columns:
We can visualize this set of two dimensional data simply by plotting it as a set of points where
is the entry in the first column and
is the entry in the second column. To simplify things we’ll consider the data to be “standardized” ahead of time. If you don’t know what that means, don’t worry about it! It just makes the calculations simpler.
Additionally, let’s highlight one data point in particular and call it as we did with the matrix representation
. This will help us see what PCA is doing to individual data points. Here’s how the graph of this data might look:
Right now, each data point is represented using the “standard basis vectors,” which are (1,0) and (0,1). This just means we can think of as
:
If we reduce the dimension of the data set to a single dimension this really means we’ll only have one basis vector with which to write not just , but all of our data. We want the “best” basis vector for the job, meaning that we can use it to get pretty close to all of the data at the same time. If, for example, we only had access to the basis vector
then the best approximation we could get of
would be
, meaning this is the closest multiple of
to
:
This is not a very good approximation of the data point . In fact every approximation of every data point using only the basis vector
just comes from “projecting” onto the x-axis, which would be really good if all of the data was already near the x-axis! However, it’s not 😦
So what would a good approximation be? We’ve already hinted at it in the previous paragraph by using the word “near.” We want the transformed data to still be near the original data, so we should minimize the total amount of distance from the original data points. At the same time we want the transformed data to fall on a straight line so that a single basis vector can approximate all of the data. This is the same as projecting the data onto the “best fit line” by definition of the best fit line! It’s the line that is closest to all of the data at the same time:
To project the data onto the best fit line we just need some basic skills picked up in linear algebra. Scale the unit vector, which we call , pointing in the same direction as the best fit line by the dot product between the data point and
,
Thus the original data point is transformed into the data point:
If we index all of the other data points:
then we can write their transformed versions in the same way:
. Again, this simply projects each data point
onto the best fit line as pictured above. Thus each of these transformed points are approximations of the original data points with the added convenience that they can now be approximated pretty well by scalar multiples of the just the unit basis vector
.
Additionally, since each of the approximations is just a scalar multiple of we can view the transformed data as just being the scalars:
themselves. Visually we can picture this in the following way:
This reveals the dual way of thinking about what this transformation of the data does. The transformed data points are now just numbers living in one dimension, and they are all quite spaced apart! In fact, they are as spaced out as possible in the following sense. If we used any other unit vector besides the one pointing in the direction of the best fit line, the resulting numbers would be less spread out. The amount of spread for a given set of numbers can be measured in several ways, but we use “variance.” Thus, by dotting our data with the unit vector pointing in the direction of the best fit line, the transformed data has maximal variance. This viewpoint is the one we take when extending the ideas into higher dimensions.
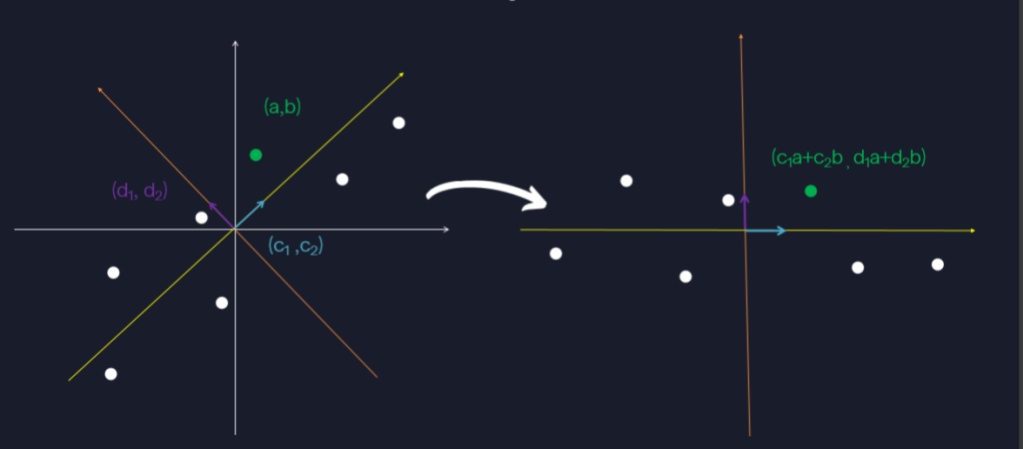
Suppose now that we do not want to reduce dimension, and instead we want to look at our data in two dimensions, but from a different perspective. This means we need a second basis vector with which to express the data besides the “best” one from before: . What should our other basis vector be? Since we still want to have a basis it makes sense to use an orthogonal unit vector obeying the same kinds of properties as the first vector.
We want the vector to still maximize variance, while at the same time being orthogonal to . Since we are only dealing with two dimensional data, this does not leave us many options, we use the vector
:
This projects the data to one dimension again, but this time by dotting with , the unit vector orthogonal to
. By turning
into both
and
we are really writing
in terms of the basis vectors
and
which we can think about visually as:
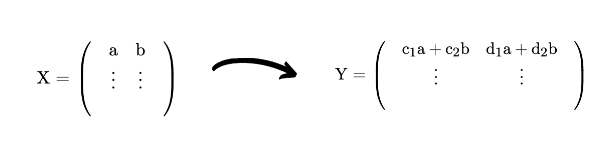
In terms of matrices we are doing:
So the simple idea behind PCA is to find these “best” vectors and
, and then dot them with the data to get transformed data. What are these “best” vectors? In two dimensions we saw that the unit vector pointing in the direction of the best fit line is the “best” because as a single basis vector, it’s the best at approximating all of the data at once by virtue of the best fit line minimizing the distance to all points. Then we noted this is equivalent to maximizing the variance of the transformed data. In higher dimensions it’s harder to visualize what’s going on, so we need to be more general and just think about if we have a matrix:
then we can project onto one dimension by doing matrix multiplication where:
and the result will be a matrix with only one column. This column is the transformed data and it’s one dimensional, so just a single column of numbers like before, written in terms of the basis vector c:
We want this resulting single column of data to have maximal variance. Recall that the variance of a bunch of numbers is given by averaging their square distance from the mean. Since we are working with standardized data the mean is 0 so the variance of is given by:
and we want this to be maximized. We can think of in a slightly different way, which you can check the by multiplying out the matrices:
The middle matrix, is an important one. It has variances of each of the columns of the original matrix X as diagonal entries. Meanwhile, the rest of the entries will be the “covariances” between any pair of columns, where in general the covariance between two standardized columns of data
and
is given by:
Name each column of ,
and the result is:
which is called the variance-covariance matrix of , or
. In general this matrix has real entries and is symmetric because
. We can now rephrase our goal. We want to find the vector c that maximizes:
Since is real symmetric we can decompose it as
where
has the “eigenvectors” of
as columns, and:
has the corresponding eigenvalues as the diagonal entries and 0’s elsewhere. An additional nice property is that is an orthogonal matrix meaning that
. Now we can show that
is automatically a unit vector because
is a unit vector:
Since and
are both unit vectors this means that
, which we are trying to maximize, must attain the same maximum as
. This follows because
and
both range over all possible unit vectors, so that the two matrices range across the same sets of possible values. This new matrix is much easier to maximize though! Note that
so that:
Now since because
is a unit vector, this means that:
can be at most where
is the largest eigenvalue of
, which occurs when
and the rest of the
. Thus:
is maximal and equals when
is the eigenvector corresponding to the largest eigenvalue,
of the variance-covariance matrix. Therefore, to summarize, if you want to transform your data:
into such that the resulting data has maximal variance:
then you should use the eigenvector:
corresponding to the largest eigenvalue of:
and this is the “first principle component,” which is really the data projected onto one dimension using PCA. If you want to project the data to two dimensions, then you need another
:
to help you get a second dimension. Following the same kind of logic, that is, wanting to maximize resulting variance, we choose to be the eigenvector corresponding to the second largest eigenvalue. This is because we want to have a basis with two vectors, so an additional constraint is that this next vector should be orthogonal to the previous vector.
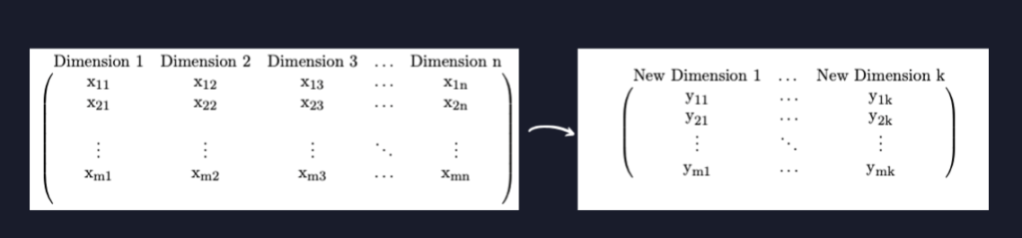
In general the result of PCA is that the matrix gets transformed into the matrix
whose columns are
where
is the ith eigenvector of
ordered by size of corresponding eigenvalue. We rename these vectors because we can’t just call them all c:
Then, to reduce the dimension of X to from dimension n to dimension k we simply keep only the first k columns of Y:
We can then think about how much of the original total variance of the data from has been retained by using another nice property of matrices. Here we think of the total variance of the data in matrix
as the sum of the variances in each of the columns of
. This is exactly the same as the sum of the diagonals of the matrix
:
The sum of the diagonals of a matrix has a name! It’s called the “trace” of the matrix. The trace happens to also equal the sum of the eigenvalues of the matrix, so we can say that the total variance of is
. If we order the eigenvalues by size then decide to keep the first
columns of the transformed matrix
this means that the variances of the columns of
will be
and so the proportion of variance retained in the columns of
is given by:
.